How is the next generation sequencing data analyzed?
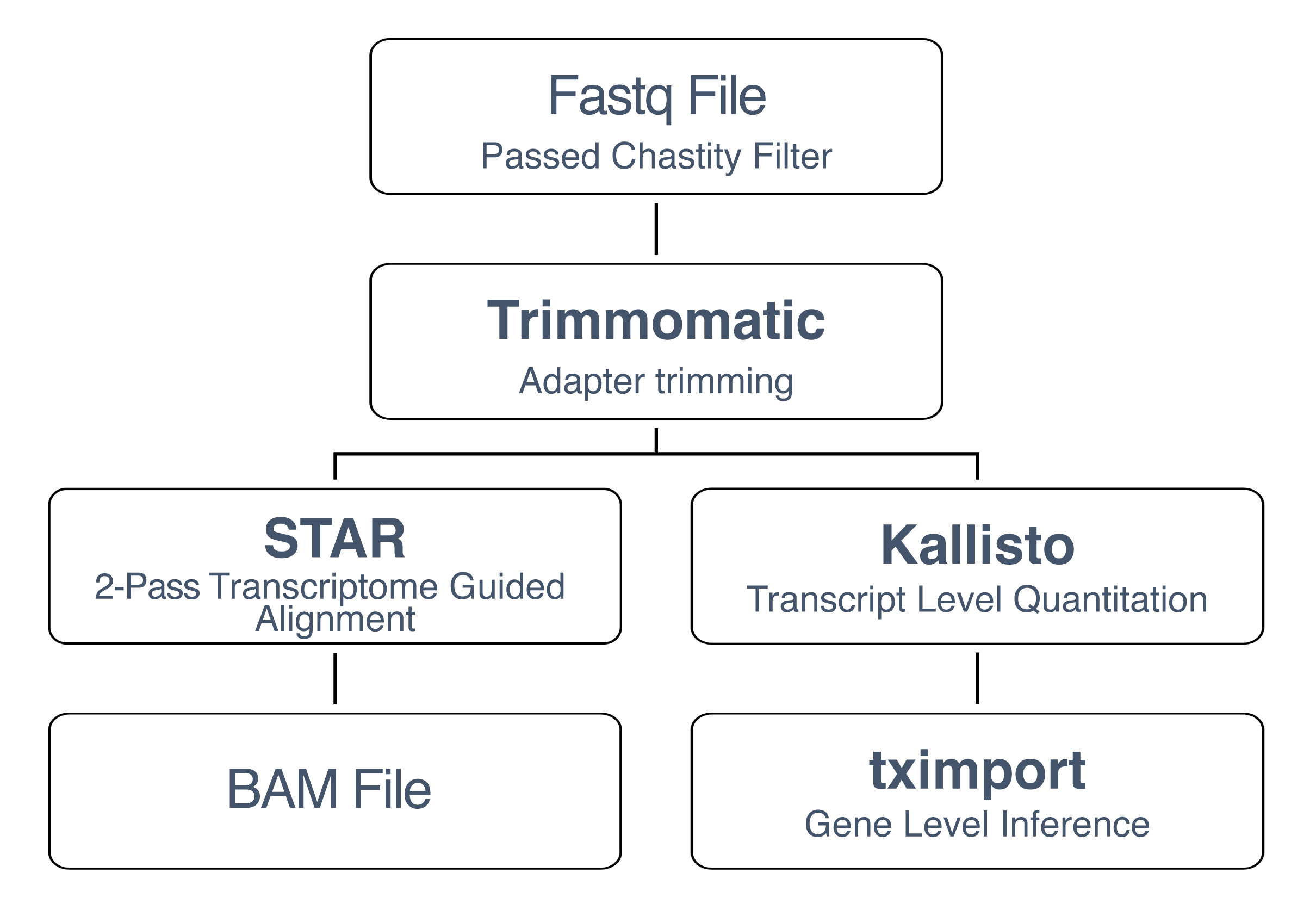
RNA-seq Data Analysis Pipeline
The analyzed data is created by taking the raw FASTQ file and processing it through the following RNA-seq data pipeline:
Ensembl DB Version
The following Ensembl database versions have been used:
- Mus Musculus - mus_musculus_core_84_38
- Homo Sapiens - homo_sapiens_core_85_38
Analysis Software
The following software is used to analyze the data:
| Software | Version & Link |
|---|---|
| Trimmomatic | v0.36 |
| STAR | v2.5.2b |
| Samtools | v1.3.1 |
| kallisto | v0.43.0 |
| tximport | v1.2.0 |
| FastQC | v0.11.5 |
PE RNAseq Snakefile
Created by Ashley Parker 11/6/14 | Last modified by Matthew Brooks 3/23/17
#######################################
# Import config file and modules needed
#######################################
import config
import glob
import os
###########################################################
# Import references needed for analysis
###########################################################
KALLISTOREF =
STARREF =
ADAPTERS =
########################################
# Import sample names from the FQ folder
########################################
SAMPLES = [os.path.basename(fname).split('.')[0] for fname in glob.glob('FQ/*fastq.gz')]
READS = ["R1", "R2"]
#############################################################
# List of directories needed and end point files for analysis
#############################################################
DIRS = ['trim/','unpaired/','unpaired/trim/','BAMS/','star/','kallisto/','fastqc/','logs/']
FQC = expand("fastqc/{sample}.{read}_fastqc.html", sample=SAMPLES, read=READS)
KAL = expand("kallisto/{sample}/abundance.h5", sample=SAMPLES)
STAR = expand("star/{sample}.star/ReadsPerGene.out.tab", sample=SAMPLES)
IDX = expand("BAMS/{sample}.bam.bai", sample=SAMPLES)
##############################
# Snakemake rules for analysis
##############################
localrules: dirs
rule all:
input: DIRS + FQC + KAL + STAR + IDX
params: time="10:00:00"
shell: "mv slurm* logs/"
rule dirs:
output: DIRS
params: time = "01:00:00"
shell: "mkdir -p "+' '.join(DIRS)
rule trimmo:
input: R1 = "FQ/{sample}.R1.fastq.gz", R2 = "FQ/{sample}.R2.fastq.gz"
output: forward = "trim/{sample}.R1.fastq.gz", reverse = "trim/{sample}.R2.fastq.gz"
threads: 24
params: mem="24G", time="01:00:00"
shell: """ \
module load trimmomatic; \
java -jar $TRIMMOJAR PE -threads 24 \
{input.R1} {input.R2} \
{output.forward} unpaired/{output.forward} \
{output.reverse} unpaired/{output.reverse} \
ILLUMINACLIP:{ADAPTERS}:2:30:10:1:TRUE \
"""
rule star:
input: R1 = "trim/{sample}.R1.fastq.gz", R2 = "trim/{sample}.R2.fastq.gz"
output: "star/{sample}.star/Aligned.out.bam", "star/{sample}.star/ReadsPerGene.out.tab", dir = "star/{sample}.star/"
threads: 12
params: mem="72G",time="4:00:00"
shell: """ \
module load STAR; \
mkdir -p {output.dir}; \
STAR --runThreadN 12 \
--genomeDir {STARREF} \
--readFilesIn {input.R1} {input.R2} \
--outFileNamePrefix {output.dir} \
--readFilesCommand zcat \
--outSAMtype BAM Unsorted \
--outSAMprimaryFlag AllBestScore \
--outReadsUnmapped Fastx \
--twopassMode Basic \
--genomeSAindexNbases 11 \
--outFilterType BySJout \
--outFilterMultimapNmax 20 \
--alignSJoverhangMin 8 \
--alignSJDBoverhangMin 1 \
--outFilterMismatchNmax 999 \
--alignIntronMin 20 \
--alignIntronMax 1000000 \
--alignMatesGapMax 1000000 \
"""
rule sort:
input: "star/{sample}.star/Aligned.out.bam"
output: "BAMS/{sample}.bam"
threads: 12
params: mem="24G", time="2:00:00"
shell: """ \
module load samtools; \
samtools sort --threads 12 {input} -o {output} -T {output} \
"""
rule index:
input: "BAMS/{sample}.bam"
output: "BAMS/{sample}.bam.bai"
params: time="2:00:00"
shell: """ \
module load samtools; \
samtools index {input} \
"""
rule kallisto:
input: R1 = "trim/{sample}.R1.fastq.gz", R2 = "trim/{sample}.R2.fastq.gz"
output: "kallisto/{sample}/abundance.h5", "kallisto/{sample}/abundance.tsv", "kallisto/{sample}/run_info.json", dir = "kallisto/{sample}"
params: mem="12G", time="01:00:00"
shell: """ \
module load kallisto; \
kallisto quant \
-i {KALLISTOREF} \
-b 30 \
-o {output.dir} \
{input.R1} {input.R2} \
"""
rule fastqc:
input: R1 = "trim/{sample}.R1.fastq.gz", R2 = "trim/{sample}.R2.fastq.gz"
output: "fastqc/{sample}.R1_fastqc.html", reverse = "fastqc/{sample}.R2_fastqc.html"
params: time="2:00:00"
shell: """ \
module load fastqc; \
fastqc -o fastqc {input.R1} {input.R2} \
"""
